Investigation Of The Potential Effects Of Host Genetics And Probiotic Treatment
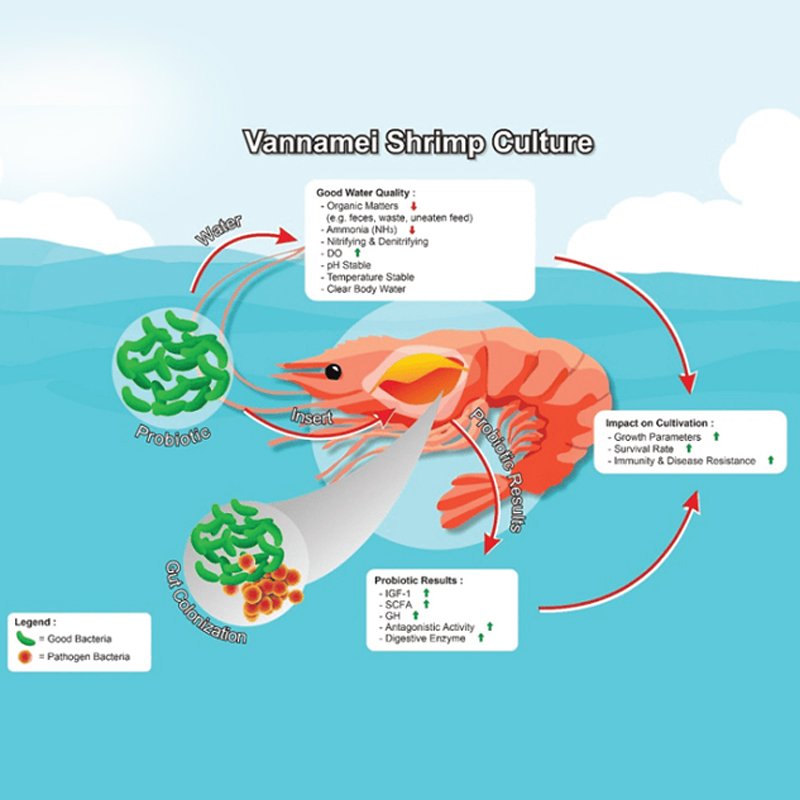
This study presents the potential effects of the genetic background and use of probiotics on the gut bacterial composition of Pacific whiteleg shrimp (Litopenaeus vannamei) grown in an indoor aquaculture facility.
The strains investigated were Shrimp Improvement Systems (SIS, Islamorada, FL, USA), a strain genetically selected for disease resistance, and an Oceanic Institute of Hawaii Pacific University (OI, Oahu, HI, USA) strain, selected for growth performance. BioWish 3P
(BiOWiSH Technologies, Cincinnati, OH, USA) was the selected probiotic.
The study consisted of two separate trials, where all shrimp were raised under standard industry conditions and fed the same diet. Shrimp were stocked in 2920 L production tanks at a density of 200/m3 and acclimated for 14 days. After the acclimation period, triplicate tanks were supplemented daily for a duration of 28 days with probiotics, while three other tanks did not receive any treatment (controls).
During the 28-day trial period, there was no statistically supported difference (p > 0.05) in either performance or health status as a result of genetic background or probiotic treatment. However, differences in gut bacterial composition, as assessed by high throughput sequencing of amplicons generated from the V1-V3 region of the bacterial 16S rRNA gene, were observed.
The relative abundance of five major operational taxonomic units (OTUs) were found to vary significantly across experimental groups (p < 0.05). Notably, operational taxonomic unit (OTU) SD_Shr-00006 was at its highest abundance in d43 SIS samples, with levels greater than d71 samples of the same genetic line or any of the OI shrimp samples.
OTUs for SD_Shr-00098 displayed a similar type of profile, but with highest abundance in the OI genetic line and lowest in the SIS shrimp. SD_Shr-00004 showed an opposite profile, with highest abundance in the SIS d71 samples and lowest in the SIS d43 samples.
Together, these results suggest that host genetic background can be an important determinant of gut bacterial composition in aquaculture-raised whiteleg shrimp and indicate that development of strategies to manipulate the microbiome of this important seafood will likely need to be customized depending on the genetic line.
Source: https://res.mdpi.com/microorganisms/microorganisms-07-00217/article_deploy/microorganisms-07-00217.pdf?filename&attachment=1&fbclid=IwAR2hK0D92p8YCW-dto4_4Nait_K08-K3nhs98hCJHXWMFrFgaRRCqUlgwY0





